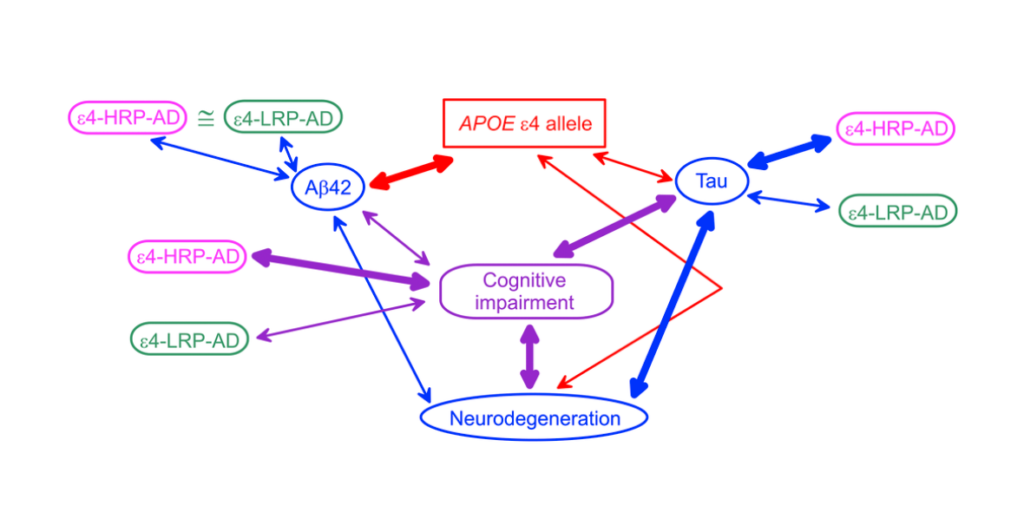
“The thickness of the arrows denotes tighter (thick lines) and weaker (thin lines) links between genetic variants, AD biomarkers, and cognitive impairment.”
BUFFALO, NY- January, 2023 – A new research paper was published on the cover of Aging (listed as “Aging (Albany NY)” by Medline/PubMed and “Aging-US” by Web of Science) Volume 14, Issue 24, entitled, “Associations of the APOE ε2 and ε4 alleles and polygenic profiles comprising APOE-TOMM40-APOC1 variants with Alzheimer’s disease biomarkers.”
Capturing the genetic architecture of Alzheimer’s disease (AD) is challenging because of the complex interplay of genetic and non-genetic factors in its etiology. It has been suggested that AD biomarkers may improve the characterization of AD pathology and its genetic architecture. Most studies have focused on connections of individual genetic variants with AD biomarkers, whereas the role of combinations of genetic variants is substantially underexplored.
In this new study for the Alzheimer’s Disease Neuroimaging Initiative, researchers, from Alexander M. Kulminski, Ethan Jain-Washburn, Elena Loiko, Yury Loika, Fan Feng, and Irina Culminskaya from Duke University and University of California examined the associations of the APOE ε2 and ε4 alleles and polygenic profiles comprising the ε4-encoding rs429358, TOMM40 rs2075650, and APOC1 rs12721046 polymorphisms with cerebrospinal fluid (CSF) and plasma amyloid β (Aβ40 and Aβ42) and tau biomarkers.
“Here, we examine the associations of the APOE ε2 and ε4 alleles and the AD-risk-differentiating compound genotypes comprising rs429358, rs2075650, and rs12721046 SNPs with Aβ40, Aβ42, and tau AD biomarkers measured in CSF and plasma using data from three studies: the AD Neuroimaging Initiative (ADNI), the Atherosclerosis Risk in Communities (ARIC) study, and the Framingham Heart Study (FHS).”
Findings from this study support associations of the ε4 alleles with both plasma and CSF Aβ42 and CSF tau, and the ε2 alleles with baseline, but not longitudinal, CSF Aβ42 measurements. The researchers found that the ε4-bearing polygenic profiles conferring higher and lower AD risks are differentially associated with tau but not Aβ42. Modulation of the effect of the ε4 alleles by TOMM40 and APOC1 variants indicates the potential genetic mechanism of differential roles of Aβ and tau in AD pathogenesis.
“Our primary finding is that the ε4-bearing polygenic profiles conferring higher and lower AD risks are differently associated with tau but not Aβ42. The other main results of our work are characterizations of the associations of the APOE ε2 and ε4 alleles with Aβ40, Aβ42, and tau biomarkers in ADNI-1, ADNI-2/GO, ARIC, and three FHS cohorts.”
DOI: https://doi.org/10.18632/aging.204384
Corresponding Author: Alexander M. Kulminski – [email protected]
Keywords: aging, apolipoprotein E polymorphism, Alzheimer’s disease, haplotypes, Alzheimer’s disease biomarkers
Sign up for free Altmetric alerts about this article: https://aging.altmetric.com/details/email_updates?id=10.18632%2Faging.204384
AGING (AGING-US) VIDEOS: YouTube | LabTube | Aging-US.com
About Aging-US:
Launched in 2009, Aging (Aging-US) publishes papers of general interest and biological significance in all fields of aging research and age-related diseases, including cancer—and now, with a special focus on COVID-19 vulnerability as an age-dependent syndrome. Topics in Aging go beyond traditional gerontology, including, but not limited to, cellular and molecular biology, human age-related diseases, pathology in model organisms, signal transduction pathways (e.g., p53, sirtuins, and PI-3K/AKT/mTOR, among others), and approaches to modulating these signaling pathways.
Please visit our website at www.Aging-US.com and connect with us:
- SoundCloud – https://soundcloud.com/Aging-Us
- Facebook – https://www.facebook.com/AgingUS/
- Twitter – https://twitter.com/AgingJrnl
- Instagram – https://www.instagram.com/agingjrnl/
- YouTube – https://www.youtube.com/agingus
- LinkedIn – https://www.linkedin.com/company/aging/
- Reddit – https://www.reddit.com/user/AgingUS
- Pinterest – https://www.pinterest.com/AgingUS/
For media inquiries, please contact [email protected].